Our Research
Histone Variant
Biology
Developmental
Enhancer Activation
Heterochromatin and
Transposable Elements
Histone Variant Biology
Our genome is packaged into chromatin, which allows cells to control the accessibility of all DNA encoded information. The selective incorporation of specialized histone proteins, or variants, into chromatin is an important feature of epigenetic regulation. A main focus of our lab is one such protein, the histone variant H3.3. The identification of mutations in H3.3 and associated proteins in human cancers and developmental disorders has heightened the need to understand the role of this histone variant in normal development and adult homeostasis.
How H3.3 contributes uniquely to chromatin function is a long-standing, unanswered question in the field. While long associated with gene activation, our recent studies establish that H3.3 also deposited at repetitive, heterochromatic regions of the genome, with deposition at each region facilitated by independent chaperone complexes.
The goals of this arm of our research program are to:
Understand how H3.3 is partitioned between its two chaperone complexes
Determine how H3.3 deposition influences local chromatin landscapes and downstream genome usage
Discover new proteins that interact with H3.3 and their role in genome regulation
Understand the functional role of H3.3 in both embryonic and adult stem cell models
Key Publications
H3.3 contributes to chromatin accessibility and transcription factor binding at promoter-proximal regulatory elements in embryonic stem cells
Tafessu A, O’Hara R, Martire S, Dube AL, Saha P, Gant VU, Banaszynski LA. (2023) Genome Biology, 24, 25. [paper]
ATRX promotes heterochromatin formation to protect cells from G-quadruplex DNA-mediated stress
Teng Y-C, Sundaresan A, O’Hara R, Gant VU, Li M, Martire S, Warshaw JN, Basu A, Banaszynski LA. (2021) Nature Communications 12, 3887. [paper]
Phosphorylation of histone H3.3 at serine 31 promotes p300 activity and enhancer acetylation
Martire S, Gogate AA, Whitmill A, Tafessu A, Nguyen J, Teng Y-C, Tastemel M, Banaszynski LA. (2019) Nature Genetics 51, 941-946. [paper]
Histone H3.3 is required for endogenous retroviral element silencing in embryonic stem cells
Elsässer SJ*, Noh K-M, Diaz N, Allis CD, Banaszynski LA.* (2015) Nature 522, 240-244. *equal contribution [paper]
Hira-dependent histone H3.3 deposition facilitates PRC2 recruitment at developmental loci in ES cells
Banaszynski LA, Wen D, Dewell S, Whitcomb SJ, Lin M, Diaz N, Elsässer SJ, Chapgier A, Goldberg AD, Canaani E, Rafii S, Zheng D, Allis CD. (2013). Hira-dependent histone H3.3 deposition facilitates PRC2 recruitment at developmental loci in ES cells. Cell 155, 107-120. [paper]
Distinct factors control histone variant H3.3 localization at specific genomic regions
Goldberg AD, Banaszynski LA*, Noh K-M*, …, Allis CD. (2010) Cell 140, 678-691. *equal contribution [paper]
Literature Review
The roles of histone variants in fine-tuning chromatin organization and function
Martire S and Banaszynski LA. (2020) Nat. Rev. Mol. Cell Bio. 21, 522-541. [paper]
Developmental Enhancer Activation
Pluripotency is a critical model for understanding fundamental principles of cell fate specification. This process makes selective use of enhancers, regulatory elements that facilitate the transcription of cell-type specific genes. Increased understanding of the molecular events that regulate enhancer activity, particularly under developmental contexts, will provide important insights into congenital diseases that occur with their dysregulation.
The chromatin state at enhancers is characterized by the presence of the histone variant, H3.3, and recruitment of the CBP/p300 family of transcriptional co-activators. Our recent studies demonstrate that H3.3 deposition at enhancers allows for variant-specific phosphorylation that stimulates p300 acetyltransferase activity towards its substrate histone H3 lysine 27 (H3K27ac) in mouse embryonic stem cells (ESCs).
The goals of this arm of our research program are to:
Determine how H3.3 phosphorylation stimulates p300 activity in ESCs
Understand the role of CBP/p300 co-activators at specific developmental enhancers
Study how developmental metabolism regulates CBP/p300 enzymatic activity
Determine how H3.3 and CBP/p300 function to promote transcription during pre-implantation development
Key Publications
H3.3 contributes to chromatin accessibility and transcription factor binding at promoter-proximal regulatory elements
Tafessu A, O’Hara R, Martire S, Dube AL, Saha P, Gant VU, Banaszynski LA. (2023) Genome Biology 24, 25. [paper]
Differential contribution of p300 and CBP to regulatory elements in ESCs
Martire S, Nguyen J, Sundaresan A, Banaszynski LA. (2020) BMC Mol. Cell Biol. 21, 55. [paper]
Phosphorylation of histone H3.3 at serine 31 promotes p300 activity and enhancer acetylation
Martire S, Gogate AA, Whitmill A, Tafessu A, Nguyen J, Teng Y-C, Tastemel M, Banaszynski LA. (2019) Nature Genetics 51, 941-946. [paper]
Histone variant H3.3 is an essential maternal factor for oocyte reprogramming
Wen D*, Banaszynski LA*, Liu Y, Xiang J, Elemento O, Rosenwaks Z, Allis CD, Rafii S. (2014) Proc. Natl. Acad. Sci. 111, 7325-7330. *equal contribution [paper]
Distinct factors control histone variant H3.3 localization at specific genomic regions
Goldberg AD, Banaszynski LA*, Noh K-M*, …, Allis CD. (2010) Cell 140, 678-691. *equal contribution [paper]
Literature Review
Establishment and function of chromatin modification at enhancers
Tafessu A and Banaszynski LA. (2020) Open Biology. 10, 200255. [paper]
Heterochromatin and
Transposable Elements
Transposable elements (TEs), including endogenous retroelements (ERVs), are widely dispersed and deeply embedded in our DNA. While these elements are silenced in somatic cells to support normal cell function, they undergo waves of programmed activation during pre-implantation development. The full complement of proteins involved in ERV regulation, and the functional consequences of ERV silencing during developmental differentiation, remain poorly understood.
Our work in this area has led to new insights into how retroelements are recognized for silencing, the impact of retroelement silencing on developmental differentiation, and how the same proteins have been used in different species to silence different retroelement sequences.
The goals of this arm of our research program are to:
Gain deep molecular insights into how different families of ERVs are silenced in different species
Identify the complete parts list of proteins involved in ERV silencing in the human genome
Explore new concepts regarding the molecular impact of ERV regulation during development and in disease
Key Publications
TASOR expression in naïve embryonic stem cells safeguards their developmental potential
Pinzon-Arteaga CA*, O’Hara R*, Mazzagati A, Ballard E, Hu Y, Pan A, Schmitz DA, Wei Y, Sakurai M, Ly P, Banaszynski LA‡, Wu J‡. (2024) Cell Reports 43, 114887. *These authors contributed equally. ‡ Co-corresponding authors [paper]
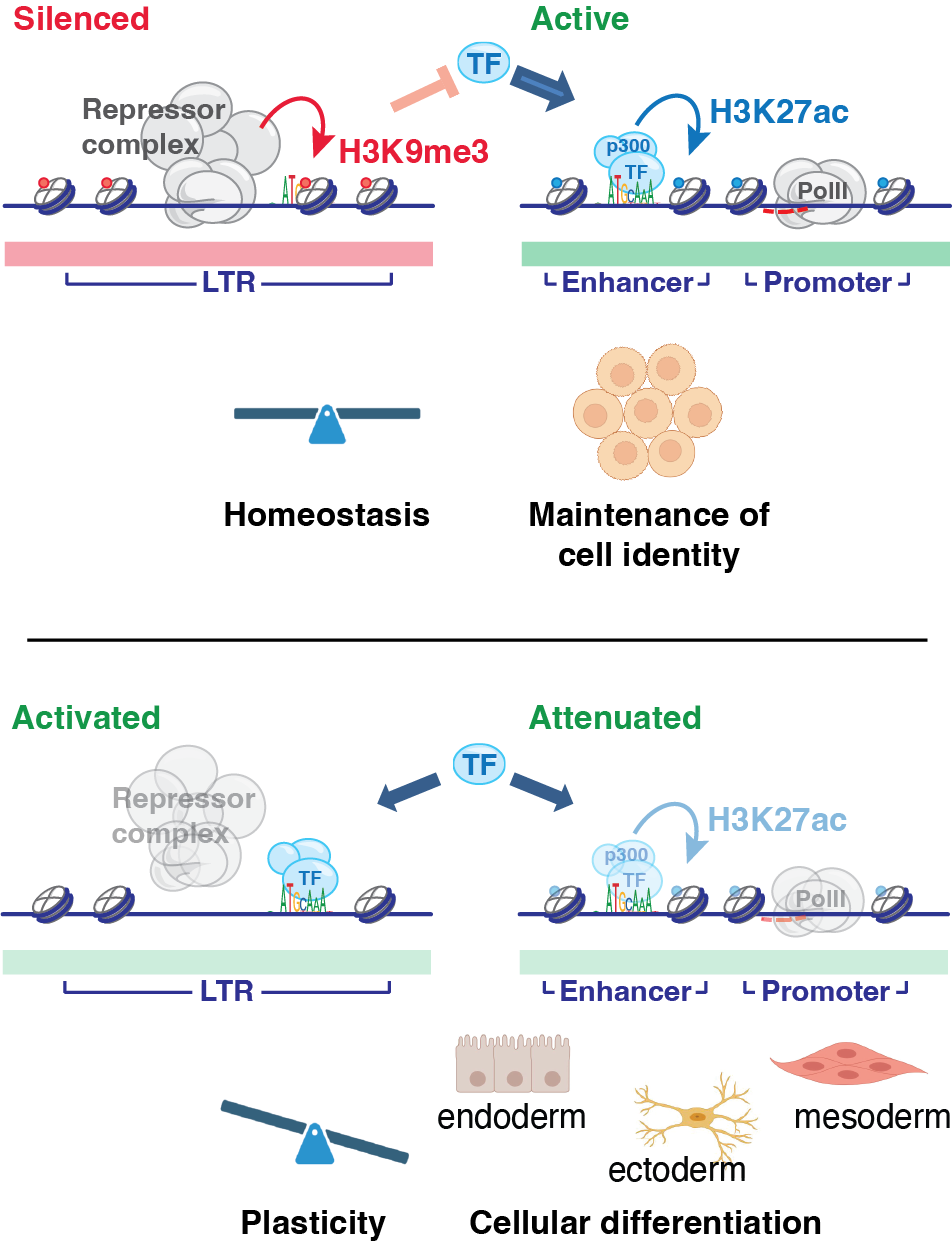
Loss of heterochromatin at endogenous retroviruses creates competition for transcription factor binding
O’Hara R and Banaszynski LA. (2022) bioRxiv. [paper]
Histone H3.3 is required for endogenous retroviral element silencing in embryonic stem cells
Elsässer SJ*, Noh K-M, Diaz N, Allis CD, Banaszynski LA.* (2015) Nature 522, 240-244. *equal contribution [paper]